Pages from the field... A Blog
|
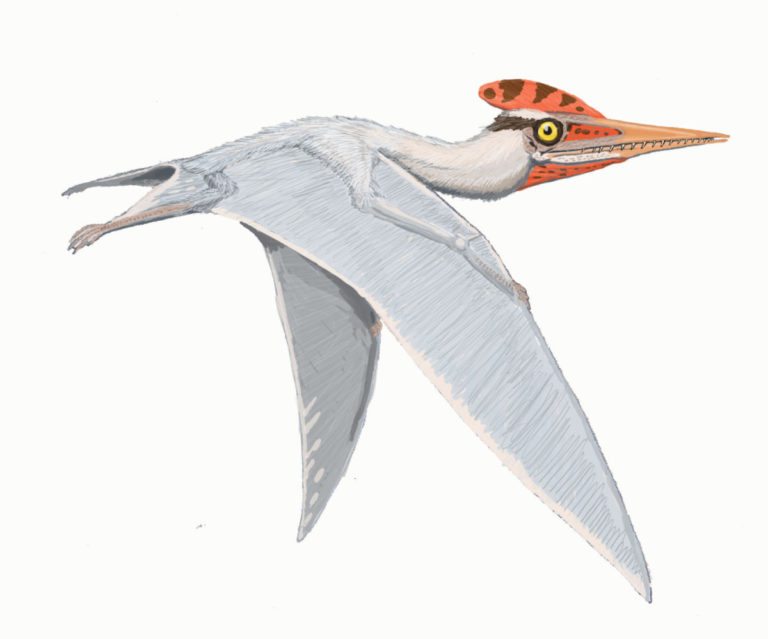
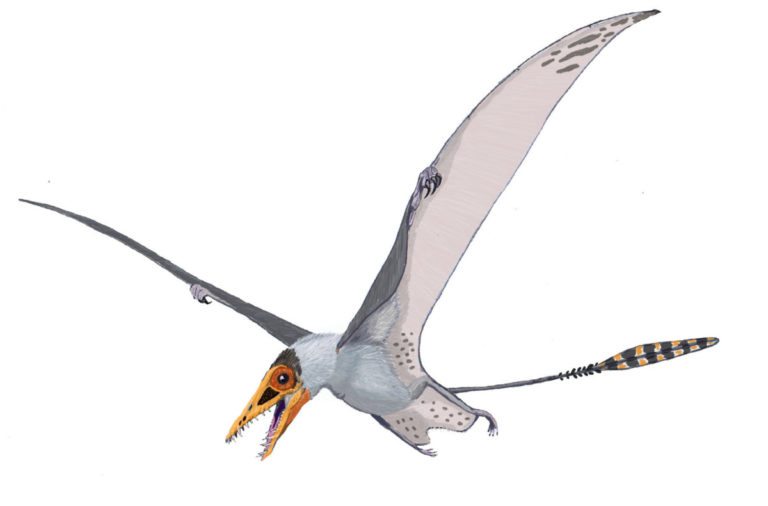
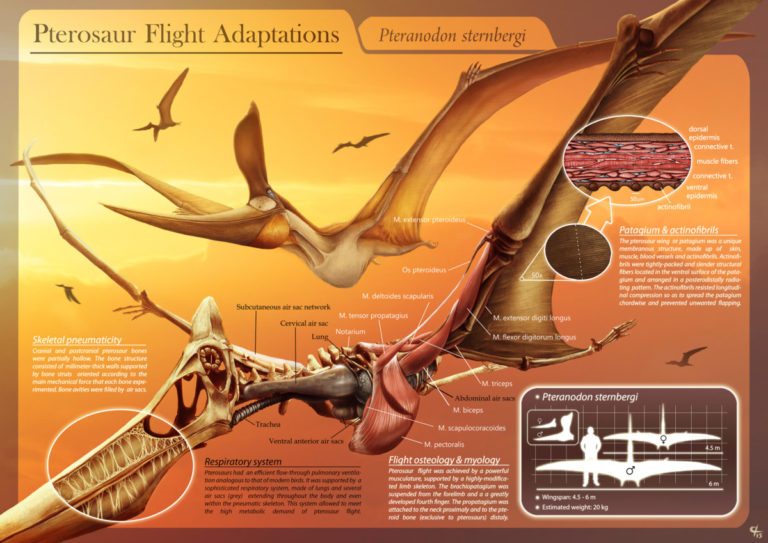
by Jack Dumbacher (First published on the Golden Gate Audubon website, With special thanks to Ilana DeBare for editing and layout suggestions!)  Eudimorphodon, pencil drawing by Arthur Weasley / Courtesy of Wikipedia Eudimorphodon, pencil drawing by Arthur Weasley / Courtesy of Wikipedia Powered flight is perhaps the most iconic ability of birds. Their feathers and wings have set them apart from other vertebrates, and they have amazing adaptations that have allowed them to master flight, including special lungs and airsacs to capture oxygen incredibly efficiently, and hollow or reduced bones and no teeth to cut down weight. There are many ways to get into the skies, and birds are not the only vertebrates that have evolved flight. Within the mammals, there are bats. The oldest known bat fossil dates to about 52 million years ago from the Green River formations in Colorado and Wyoming. Although bats are less efficient flyers than birds, bats are more maneuverable. Their wings consist of a thin membrane of skin called the patagium, stretched between their four finger bones and the side of their body. Because their fingers are so long and spread through the patagium, they have superb control of the shape and size of their wings, making them incredibly agile fliers. But birds have been flying for nearly three times as long as bats. The oldest bird fossils are from Archaeopteryx, found in the Solnhofen limestone quarries of Bavaria, Germany, and these fossils were dated to about 150 million years ago. These Archaeopteryx fossil bones mark the beginning of birds in the fossil record. Most of the close dinosaur relatives of Archaeopteryx were covered in feathers, and some even had wings. We call this group of dinosaurs “Paraves,” and they include some four-winged dinosaurs (like Microraptor, having feathered forelimbs as well as hindlimbs) and many others that were fierce terrestrial carnivores, like the Velociraptor made famous by Jurassic Park. (The film incorrectly shows Velociraptor with scaly skin instead of feathers!) Representatives of the most bizarre and largest flying vertebrates were also around at the time of Archaeopteryx. In fact, the first fossil of this group to be described was collected from the same Solnhofen limestone formation of Bavaria that produced the famed Archaeopteryx specimens. That fossil was described in 1784, and is now known as Pterodactylus antiquus, which derives from Latin for “ancient wing finger.” This fossil creature became known as the pterodactyl, and it is just one species of a huge group of flying reptiles called the Pterosaurs. Pterosaurs were capable of powered flight, and species from this group of reptiles lived from 228 to 66 million years ago, dying out along with the dinosaurs at the end of the Cretaceous. In many ways, their evolution is mirrored by that of birds. For example, pterosaurs started out with mouths full of teeth, but over time evolved a lighter, toothless, beak-like mouth. They also evolved a lung-airsac system similar to but predating that of birds by some 70 million years, that allowed efficient oxygen exchange for powered flight. Evidence from their lifestyle and body covering suggests that they were warm blooded as well. They are also different from birds in many respects. First and foremost, the pterosaurs were not part of the tree of life called Dinosaurimorpha (which gave rise to the dinosaurs), and so the pterosaurs were NOT dinosaurs. They had no feathers, although at least some did have hair-like filaments that scientists call pycnofibers. And their flight is poorly understood. As in bats, a membrane extends from the forewing and finger bones to provide shape and structure to the open wing, but the pterosaurs use only a single finger at the front of the wing to support the wing. The patagium, or wing membrane, of the pterosaurs was thicker, stronger, and more complex than that of bats. The pterosaur patagium included layers of muscle fibers as well as connective tissue called “actinofibers” that may have provided strength and support, and helped shape the wing like the battens on a sail. Many of the pterosaurs also evolved strange crests and head shapes, whose purpose is not understood. Pterosaurs were the largest animals ever known to fly. Although many pterosaurs were small and in the range of modern bird sizes, several had wingspans that exceeded five meters (16 feet). In contrast, the largest flying bird was the extinct Argentavis magnificens, or the Giant Teratorn, which had a wingspan of 5-6 meters (17-20 feet) and stood about 1.5-2 meter (5-6 feet) tall. Although this is extremely large for a bird, the are dwarfed by the larger pterosaurs. The largest of the pterosaurs was Quetzalcoatlus northropi, which had a wingspan of 10-11 meters (up to 36 feet!) and stood 4-5 meters tall (16 ft). With such large bodies and wings, they are believed to have flown at fast speeds (up to 75 miles per hour) and could cover large distances. Based upon fossil footprints from pterosaurs, they were believed to walk on four legs, with the wings folded back along the arm, and the other fingers opened out to form a hand or paw. This would have been useful for supporting their large bodies while on the ground. So pterosaurs weren't birds (and not even dinosaurs), and pterosaurs existed on earth longer than birds (although birds are still living, so they may break the Pterosaur's record). In many ways pterosaurs and birds were both moulded and engineered for flight, and the fossil record of both groups shows interesting similarities in their evolutionary trajectories.
Right now, the California Academy of Sciences has a traveling exhibit featuring pterosaurs, so you can see some real pterosaur fossils, life-sized models of pterosaurs, and learn about their biology. The exhibit was produced by the American Museum of Natural History and will remain on site through January 7, 2018.
0 Comments
In late 2010, two fascinating articles appeared that described a new and very serious avian disease (1,2). So-called "avian keratin disorder," or AKD, caused birds' bills to radically overgrow. With these outsized bills, birds were unable to preen or even properly feed -- and in their Alaskan environment, where birds were first found with the disorder, this spelled almost certain death by starvation and hypothermia. The second article suggested that the disease wasn't limited to just a couple species, and also showed that it appeared to be spreading to both more birds and to more localities. The disorder was found in chickadees, nuthatches, crows, jays, woodpeckers, hawks - and these are just the easy-to-see birds that tend to hang out at people's feeders or urban parks. The articles were partially a call to arms and a plea for more information from citizen birders and other scientists to recognize and help track the disease. Despite investigating several possible causes, the authors were unable to determine what caused the beak deformities. They looked at known viruses, bacterial infections, fungus, mites - even potential environmental toxins. But no smoking gun.
At the same time, I was starting a collaboration with Joe Derisi, who is a virus researcher at University of California San Francisco and clever virus hunter (among other things). He had some fancy lab equipment and techniques for finding viruses. One tool was a virus microarray chip -- their virochip -- a small glass microscope slide with tens of thousands of short sequences that are made to match a portion of the genome of virtually every known virus. By putting DNA from a sick person (or bird) onto the microarray, a DNA match would light up and indicate a virus present, and potentially which one. We thought it would be fun to see if his virochip could identify a bird virus. So I quickly contact the key investigators, Colleen Handel and Caroline van Hemert from the USGS labs in Alaska. They were keen to try anything, and consented to send some samples taken from sick birds. We tried the virochip, and got some hits that we followed up, but answers weren't quickly forthcoming. One challenge was that bird DNA was very different from human DNA, so we didn't know what the background virochip pattern would look like when normal bird DNA was run on the chip. But it did suggest some leads that we followed up - one that led to the discovery of a new tiny circular virus (3). But after following up on this new virus, we decided that it probably wasn't related to the disease. But by then, we were hooked and invested in finding the cause. Derisi lab members were experimenting with another technique that we wanted to try. They were using new whole-genome sequencing techniques to look for viruses. It worked something like this: Take a sample of blood or a mouth or cloacal swab from a sick bird, then extract DNA and RNA from the sample. Then shear up the DNA and RNA into billions of fragments about 400-500 bases long. Then use new Illumina genome analyzers to read the sequences of literally hundreds of millions of these fragments. Then use a computer to look for pieces that match (or nearly match) any known virus genomes. Finally, starting with a piece that matches a virus, try to build longer and longer sequences by searching the sequence dataset for additional sequences that overlap the starting sequence, and then continue extending as far as possible. Once you get these extended pieces of DNA sequence, some will be big enough to reliably determine what they match -- such as the bird's genome, or a viral genome, or bacterial genome, and so on. Around this point, we got a post doctoral researcher, Maxine Zylberberg, to help with the virus work, and we set to work in the lab. We got additional Alaskan samples from Colleen and Caroline, and Maxine also added samples from some sick hawks from California. A team from Illumina generously offered to provide an entire flow cell of sequence data (amounting to nearly a billion short sequences, worth several thousand dollars at the time!!!) for the project. Before long, Maxine and Joe and I were looking through gene fragment assemblies that Maxine had produced, and two of the fragments provided a hit to a distantly-related picornavirus known from turkeys. Although far fewer than 50% of the amino acids were identical, the new sequence could be classified as a picornavirus by the genome structure, gene order and content. If you don't speak virus, this all just means that this new virus was very distantly-related to any known virus, but it was a virus for sure. And Maxine used some other cool sequencing techniques to sequence the rest of the virus genome from the bird samples, finally nailing the ID of this new virus. What we didn't know at the time was that identifying this virus was the easy part. Maxine then worked for a couple years trying to definitively link this virus to the disease. She was able to screen lots of samples - and the birds with the disease usually tested positive for the virus (19 out of 19 Black-capped Chickadees), and healthy birds usually tested negative (7 out of 9 healthy chickadees were negative.) But this isn't enough to prove that the virus CAUSED the disease. She tried to culture the virus in the lab, which turned out to be far from easy. And without cultures, it was impossible to do so-called challenge experiments where healthy birds are exposed to the culture to see if the disease can be spread. She additionally screen other bird species, and she was able to test various tissues to see where the virus hangs out. She named the new virus "Poecivirus" after the Black-capped Chickadee (Poecile atricapillus) that it came from. And Maxine did a lot of other cool stuff - all of which is reported in our recent paper (4). During the time we were doing this work, cases of AKD were documented throughout Canada and the lower 48 states, and even a couple cases were documented in UK and Europe. The disease continues to spread in Alaska. Even if this new virus is the cause of the disease, we do not know whether it is spread directly from bird to bird, or through the environment by contaminated water or food (like bird feeders) or whether it is spread by a vector such as mosquitos. There is still much to learn about how serious the threat is to birds worldwide and how quickly it is spreading. So now, with a viable candidate for the cause of AKD - what is next? There is a role for every birder out there - especially those who take photos. You can submit information about cases and photos of sick birds to USGS at their AKD website, or directly on their reporting site. Alternatively, you can email photos and information directly via this email link. This will be most helpful as Colleen and her team continue to track the spread and severity of the AKD outbreak. Bibliography:
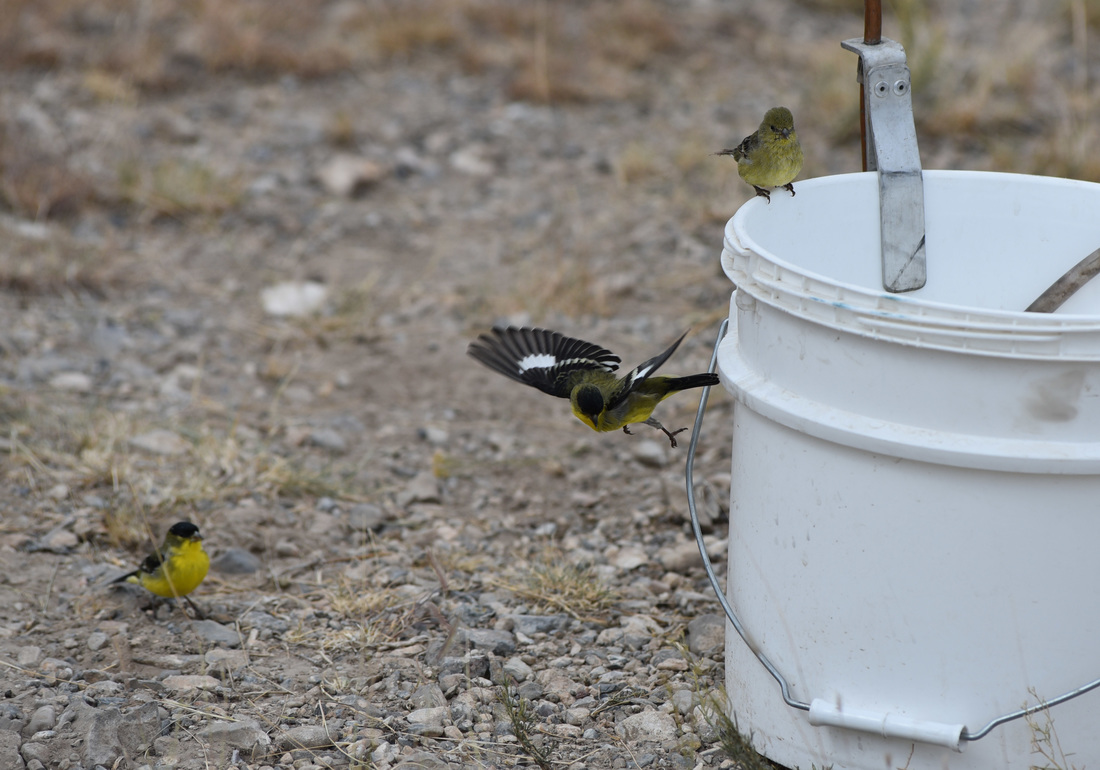
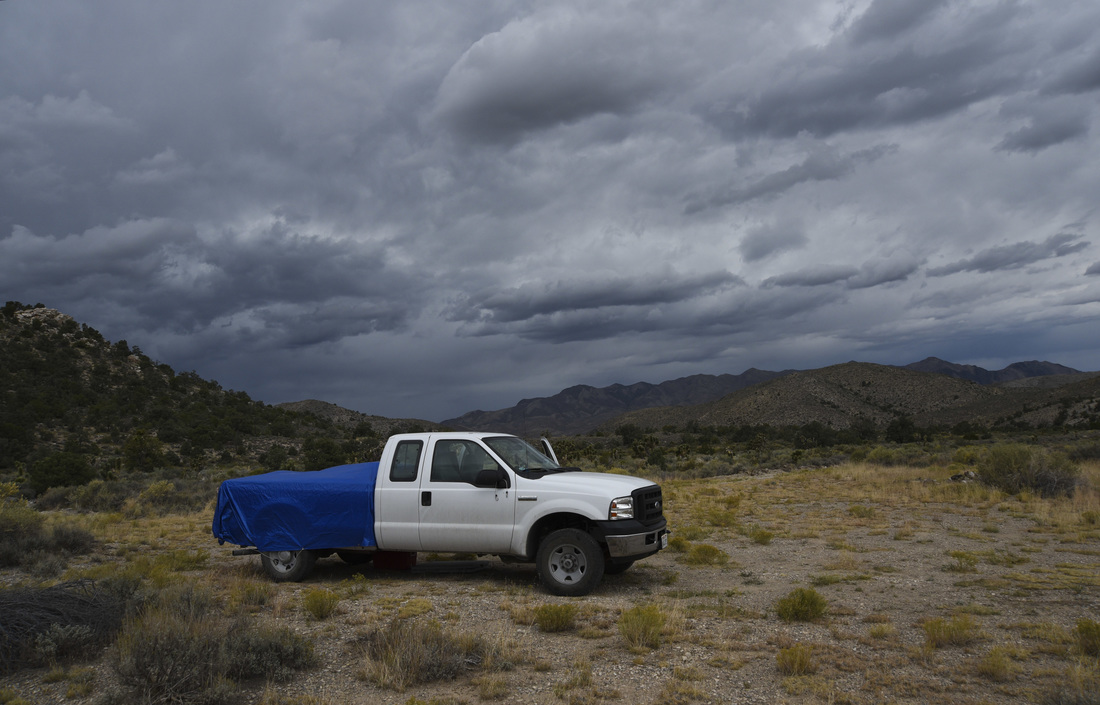
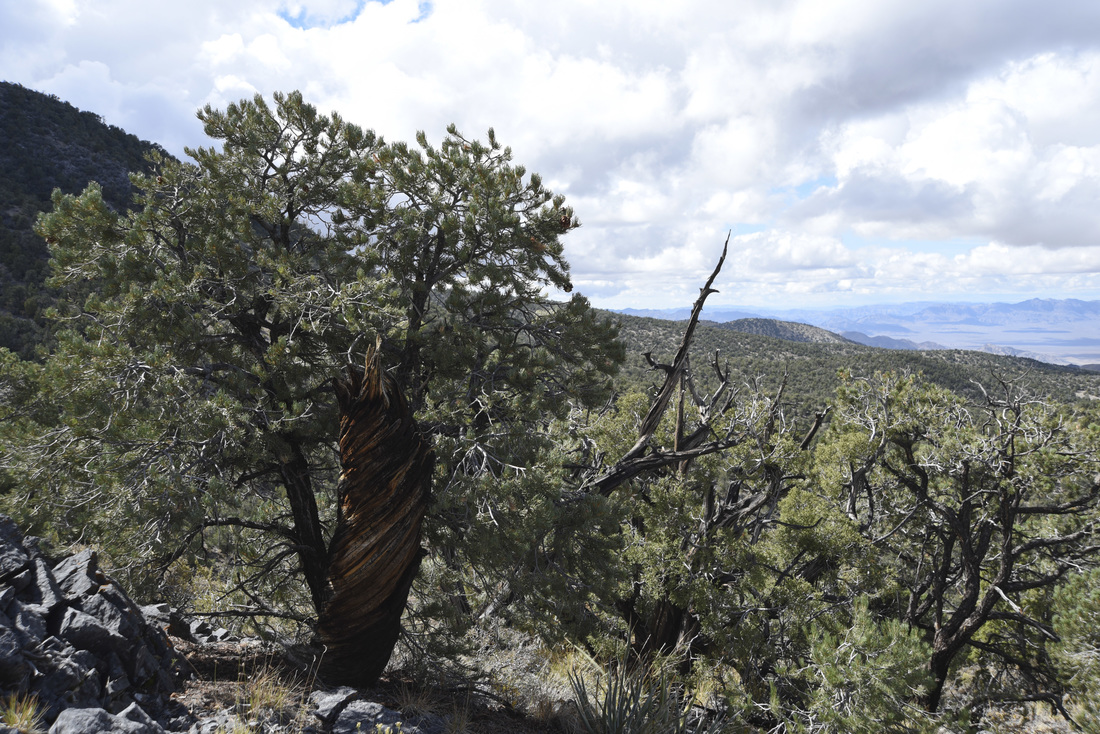
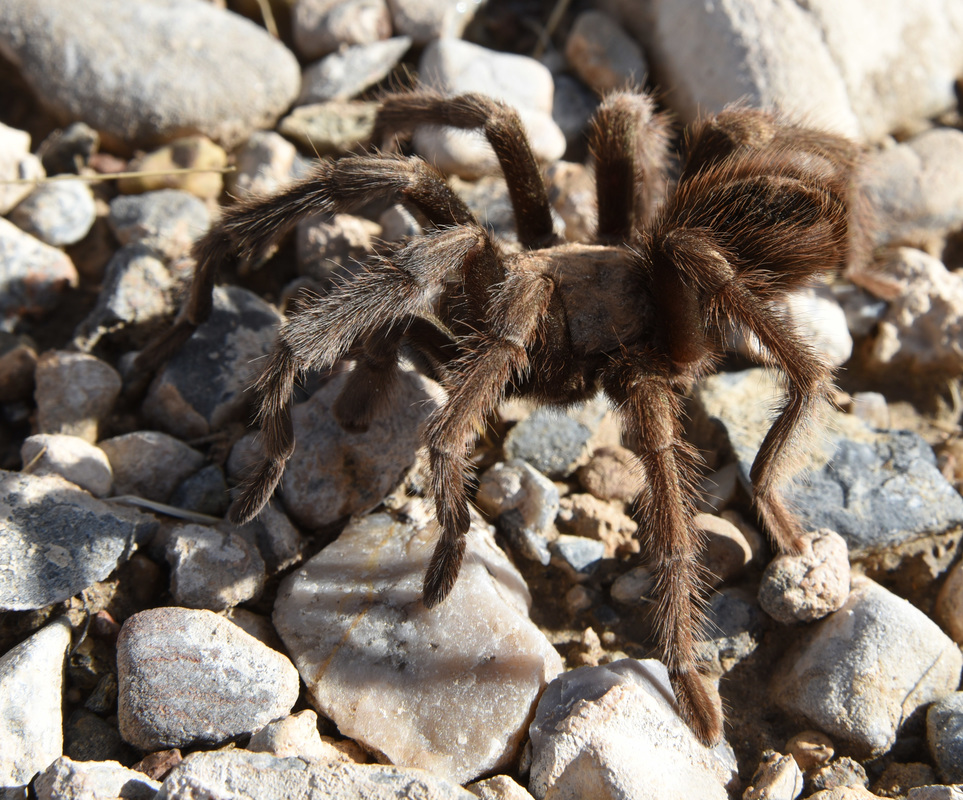
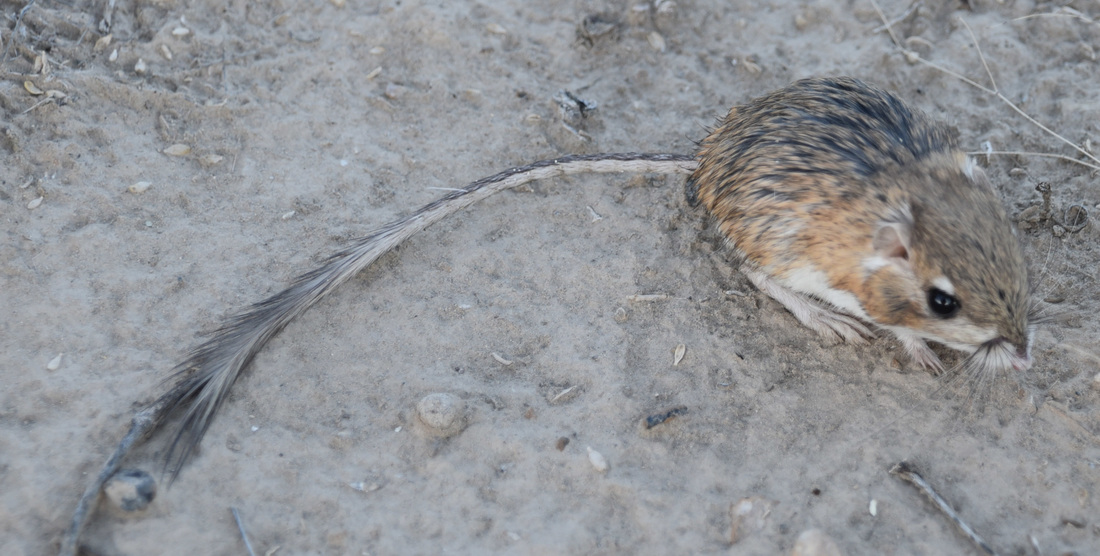
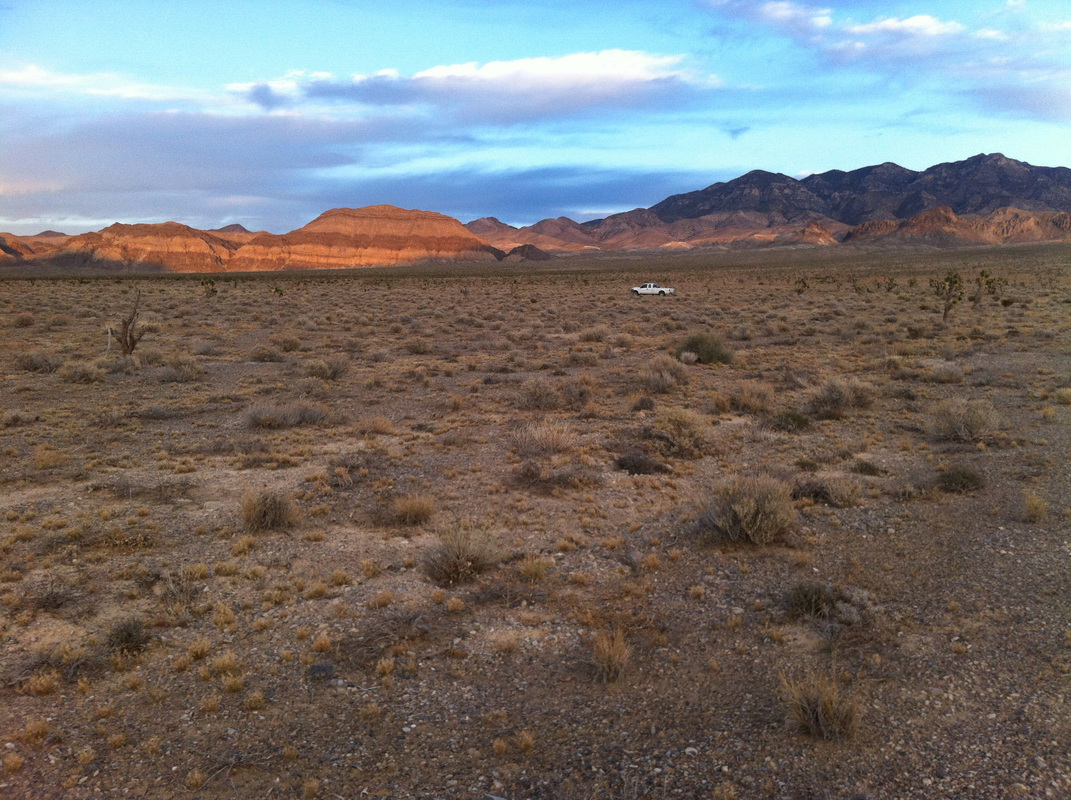
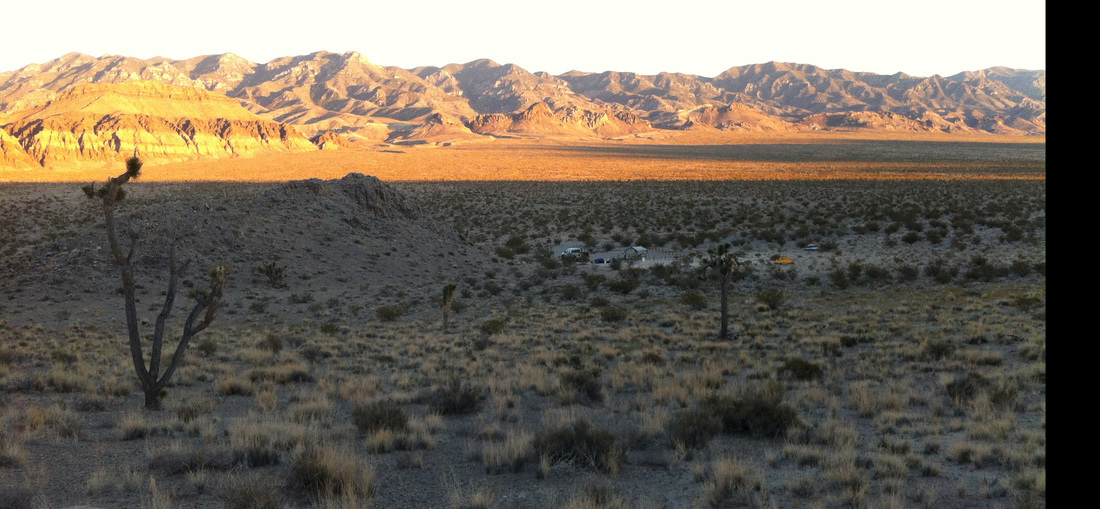
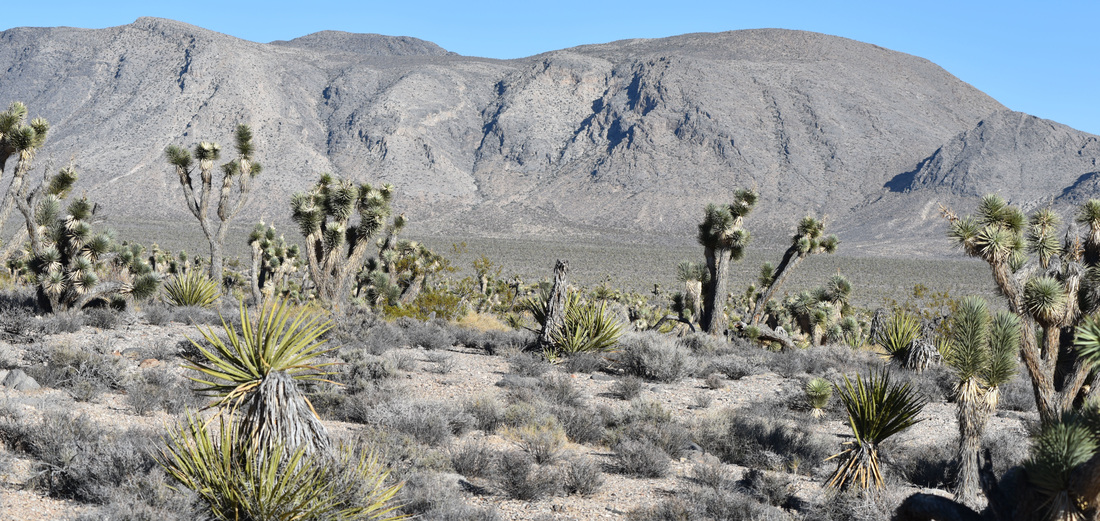
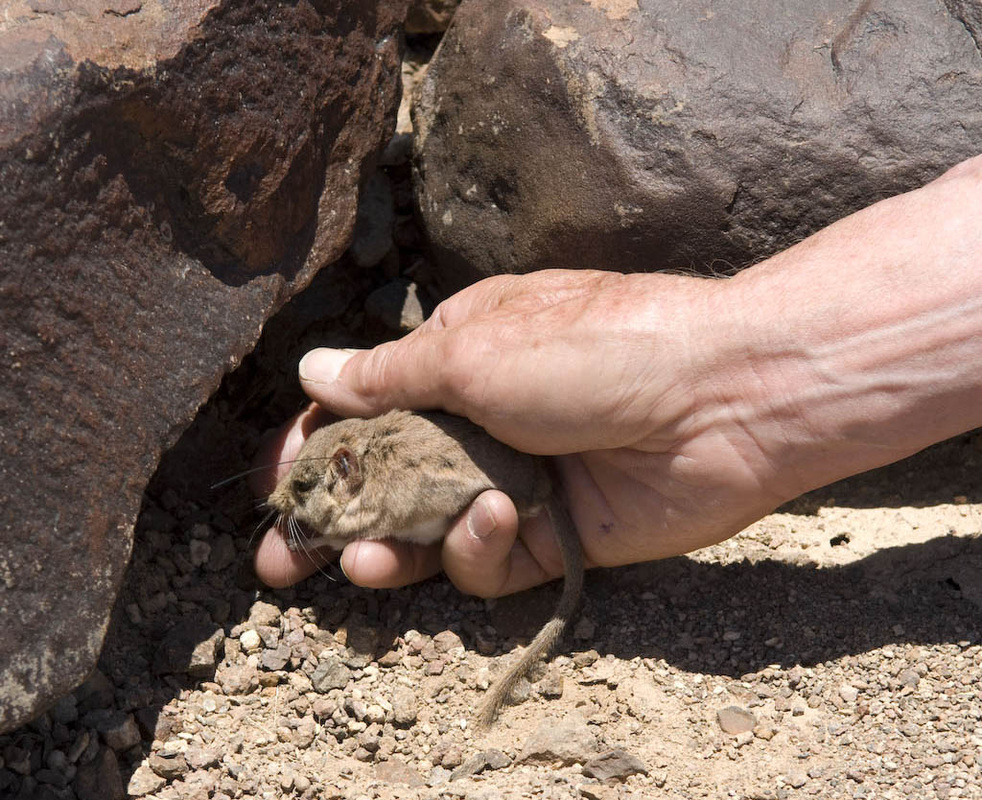
Also - Click here to download our California Academy Press release, with more information. And check out this article about the work, on the Wildlife Society Blog. I just wrote this blog about how rapidly birds can see, based upon some cool new work on bird vision. But it is posted on the Golden Gate Audubon Society Blog - so it is best to read it there. Here is a direct link... Enjoy! The day after my last post, we drove up Alamo Road to Dead Horse Road, which follows a difficult wash from the basin west of the Sheep Mountains, up around Dead Horse Ridge and into the pinyon pine and juniper forests of the Sheep Mountains. It took us a long time to drive the "road", with a few stops for moving large rocks and checking to make sure that we were on the right road. At the end of the road, we set up our camp and set out about 100 small rodent traps to see what was lurking in the brush. The temperature was cool, and we could feel the weather changing, and even the birds seemed antsy - a couple flew right into our camp and seemed very unafraid of us. Lesser Goldfinches playing in our wash water... We had lots of birds around the camp - including Pinyon Jays (50+), Northern Flicker, White-crowned Sparrows, Chipping Sparrows, Vespers Sparrows, Cassin's Finches, and of course Red-tailed Hawks, and Common Ravens. On our second night, the sky darkened, the wind picked up, and some serious clouds rolled in. Because we were a long way up a seriously-difficult-to-drive wash, we had concerns about the road washing out. Although we were ready to do what we needed to do to get out, we were lucky that the storm didn't bring much overall rain, and spread it over more than 24 hours - giving the ground plenty of time to soak up the water.  And of course - the end of the rain was mixed with bouts of bright sun and beautiful rainbows. Nothing like a rainbow in the desert... When the rain lifted, we took a long walk up to Perkin's Spring, to check out the condition of the drinker and to look for chipmunks that might be unusual. The views (like the view above) was beautiful from the slopes of the Sheep Range (looking west here toward Alamo Road). Although we saw many cliff chipmunks, we didn't see or hear any chipmunks that resembled the threatened Hidden Forest Chipmunk. Back in camp, we saw a few tarantulas, probably the desert tarantula (Aphonopelma iodius) that is common in the area. The males are commonly seen making long treks in the evening light - often walking along roads or walks that remain warm from the afternoon sun. This is Merriam's kangaroo rat (Dipodomys merriami), the most common k-rat in the area. We caught these at every elevation where we trapped - from Dead Horse Road camp to White Sage Flat. Although I tried to get photos of several other small mammal species, they all moved too fast when I let them go to get good photos. Even this guy was a little out of focus. After several days of working and trapping along Dead Horse Road, we came back to the basin and White Sage Flat. The above photo was taken from White Sage Flat looking back at the car on Alamo Road and the Sheep Ranges in the background. Very few Joshua trees lived on the flat, although creosote and J-trees were common in the habitats all around the old dry lake bed to the west of Alamo Road. Here you can see our camp just north of White Sage Flat. We trapped up into the hills and around the camp on our last night (in addition to trapping some of the northern end of the old dry lakebed) but pretty much all we caught were Merriam's kangaroo rats, some Peromyscus deer mice, and some Onychomys grasshopper mice. In fact - the grasshopper mice were quite common here - named after their food, since they eat largely insects and other arthropods, unlike most mice that are granivorous. I have heard that Onychomys is resistant to scorpion venoms, because they have co-evolved with scorpions, and they can resist even the most toxic species of Centruroides scorpion that could be dangerous to humans.
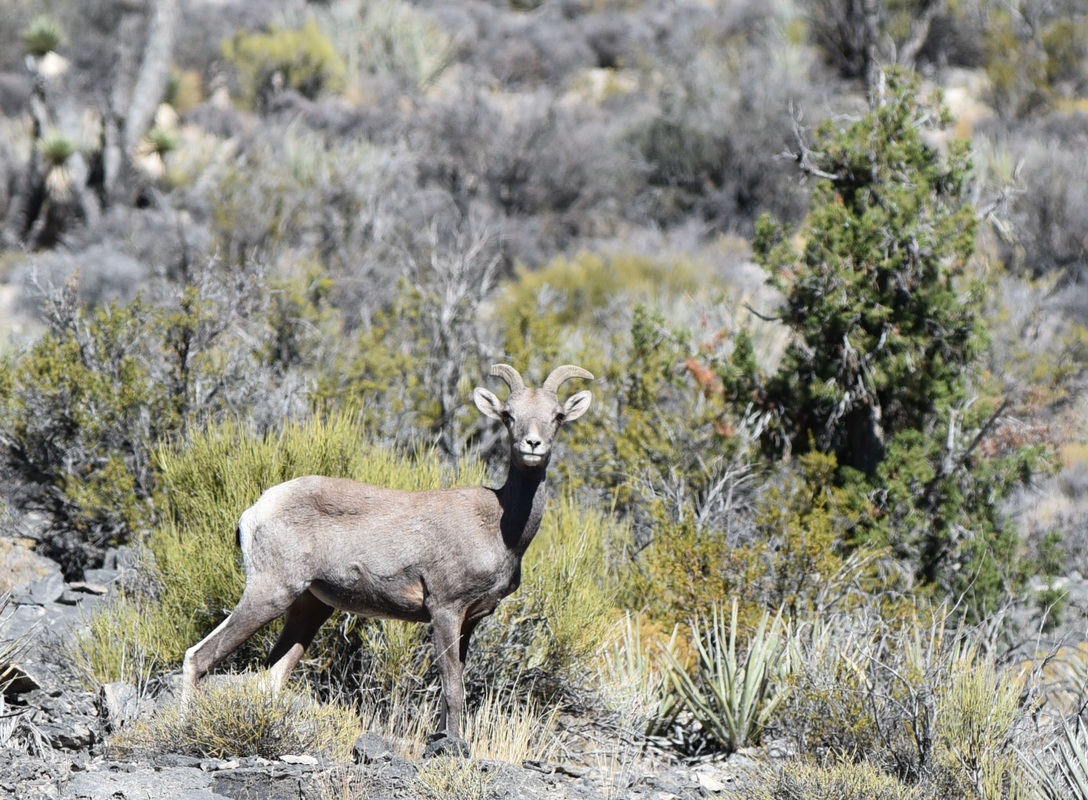
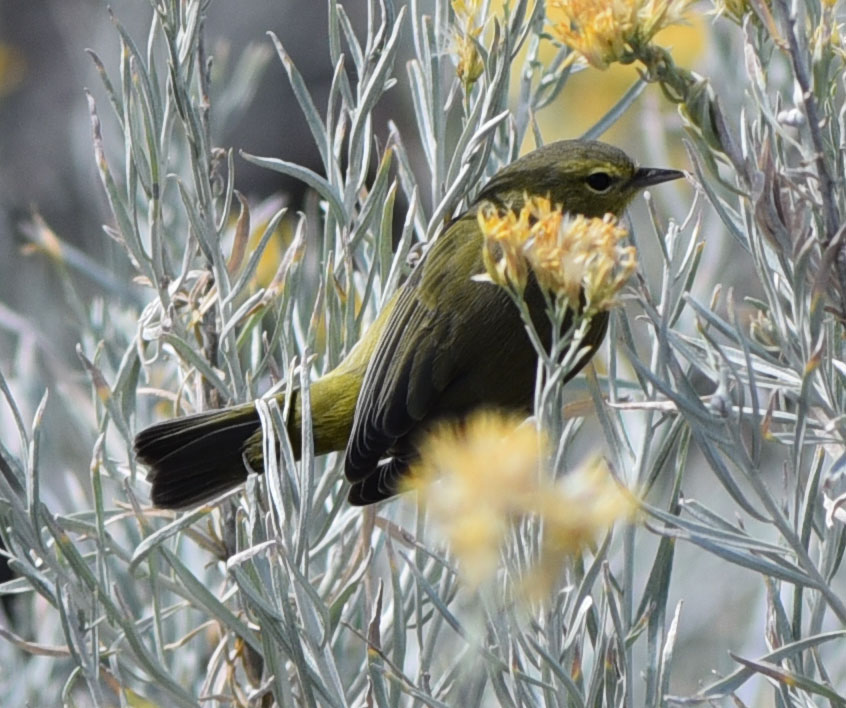
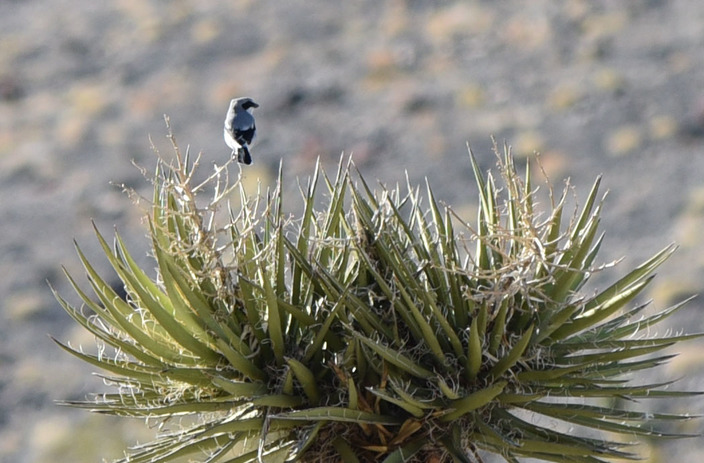
Overall, the trip was great, and we got excellent data from a variety of small mammal species. The list of small mammal species that we documented on this trip included the following: Desert Cottontail (Sylvilagus audubonii) Black-tailed jackrabbit (Lepus californicus) White-tailed antelope ground squirrel (Ammospermophilus leucrurus) Long-tailed pocket mouse (Chaetodipus formosus) Merriam's kangaroo rat (Dipodomys merriami) Chisel-toothed kangaroo rat (Dipodomys microps) Desert woodrat (Neotoma lepida) Southern grasshopper mouse (Onychomys torridus) Western harvest mouse (Reithrodontomys megalotis) North American deermouse (Peromyscus maniculatus) Canyon deermouse (Peromyscus crinitus) Cactus deermouse (Peromyscus eremicus) Pinyon deermouse (Peromyscus truei) several bats that we could not ID Coyotes (Canis latrans) calling at night (but never seen) Desert bighorn sheep (Ovis canadensis) And here is a list of bird species that we documented: Turkey Vulture Northern Harrier Cooper's Hawk Red-tailed Hawk Sandhill Crane Great Horned Owl Common Poorwill White-throated Swift Belted Kingfisher Red-naped Sapsucker Ladder-backed Woodpecker American Kestrel Prairie Falcon Say's Phoebe Loggerhead Shrike Pinyon Jay Western Scrub-Jay Common Raven Mountain Chickadee Bushtit Red-breasted Nuthatch Rock Wren Cactus Wren Ruby-crowned Kinglet Sage Thrasher American Pipit Cedar Waxwing Chipping Sparrow Brewer's Sparrow Dark-eyed Junco White-crowned Sparrow Sagebrush Sparrow Vesper Sparrow Song Sparrow Spotted Towhee Red-winged Blackbird Western Meadowlark Great-tailed Grackle House Finch Purple Finch Cassin's Finch Pine Siskin Lesser Goldfinch Lawrence's Goldfinch We're back at Desert National Wildlife Refuge, for another year of mammal surveys. But the birding was amazing today too... If you don't know where Desert NWR is, it is just north of Las Vegas - the major landform that you can see looming over the city as you drive up highway 95, buzzing by the casinos and bright lights. But just past the city is the largest national wildlife refuge in the lower 48 states, and it is in beautiful shape. They have a new visitor center that is very nice, and right by Corn Creek Spring - an amazing little natural oasis in the desert, and now a brand new PAVED road to Corn Creek from highway 95. We received a small contract from US Fish and Wildlife to help survey the natural resources - which for us is helping survey the birds and mammals. We started today up Mormon Well Road and driving to Desert Pass, and some highlights included seeing a big buck mule deer at the spring, and a desert bighorn sheep ewe in Peekaboo Canyon. Lots of birds too - we saw over 25 species while driving (which is a good number for that drive) and the desert is beautiful. Here are a couple photos, but I will blog more next time we are back at Corn Creek and have some cell reception! The bighorn sheep had a broken right horn. Unlike antlers, which grow new each year, this horn will remain broken for its entire life. Still a very handsome sheep! Orange-crowned Warbler just above Mormon Well. Bird life here was quite busy - lots of birds coming to drink from the spring and feed on the lush vegetation. Loggerhead Shrike atop a Joshua tree near Fossil Ridge...
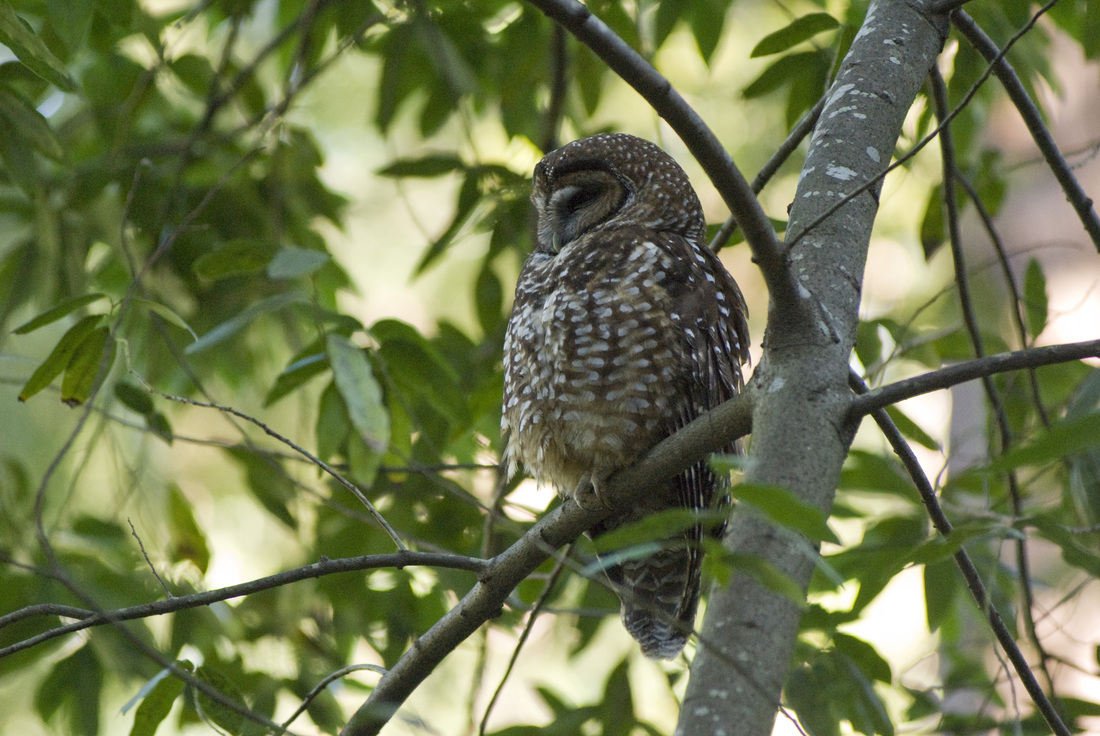
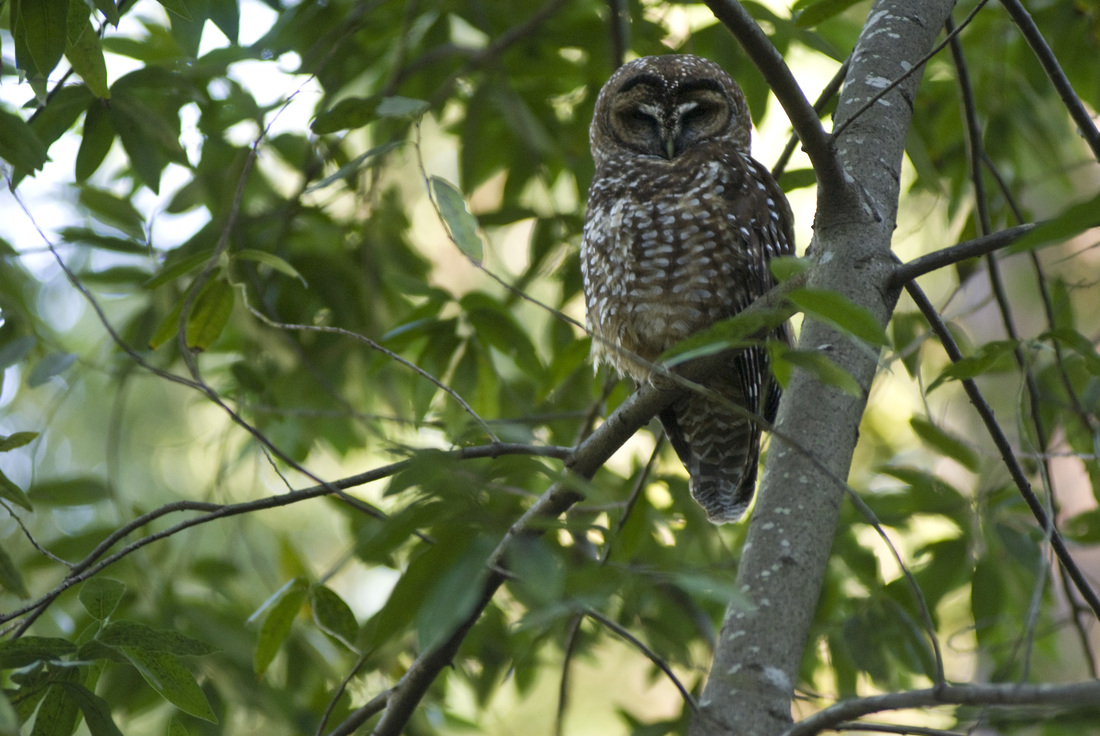
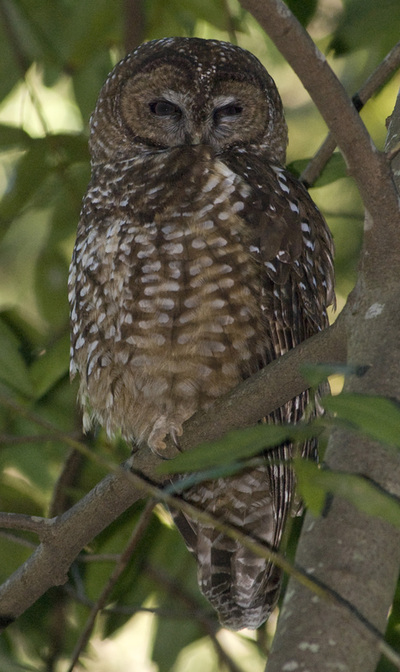
It is wonderful to live so close to San Francisco and yet still have open space adjacent to our neighborhood. We have several redwood trees in the yard, and a trailhead basically just past our house. The neighbors with dogs pass by regularly, and we get to know many of the neighbors by name. Yesterday one of our neighbors came up to the house to tell us about a Northern Spotted Owl sitting in a tree just up the road. I've heard the owls nightly for the past week or two, hooting their four-note contact calls. They've nested just up the canyon for over a decade, and they appear to be working on another nest this year, although I don't know exactly where they are building. But when I heard that owl was easy to see, I grabbed my camera and took a short walk with my wife and little girl. Although we hear the owls often, it is unusual to get a nice opportunity to see them. Northern Spotted Owls are secretive and don't come out much during the day. They blend in with the habitat well, and so long as they stay motionless, they can be very difficult to pick out amongst the trees. Many crows live in the area too, and if anything betrays the owl's location, the crows will mob and hassle them endlessly until nightfall. So although he was awake, and only about 20 feet above the ground, he was staying very still and trying not to bring any attention to himself. But his glance did follow the hikers that passed by - especially those with little dogs.
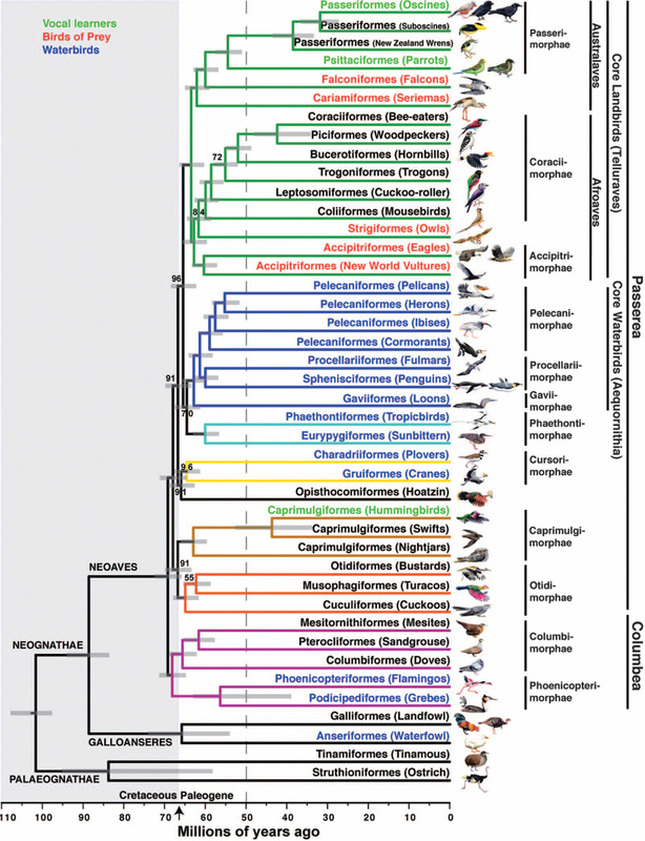
About a month ago, I recorded the owls doing a somewhat unusual call, and emailed my colleague and owl expert, Lowell Diller, who lives and works in Humboldt County. This call is quiet and a bit upslope from me, but the recording is here if you want to listen to it. Lowell had heard this call before, and suggested that the owls were establishing their central core, and might have interacted with an interloper - perhaps another Spotted Owl, or perhaps some other threat to their territory. Whatever the threat, this was definitely an aggressive call and suggested that they were probably nesting and had something worth defending. So I will keep and ear and eye out for the nest and for youngsters this year!  A recent issue of Science magazine published a suite of amazing papers based upon full genomes sequenced from 48 bird species (and a few close relatives.) They not only provide an amazingly complete and well-resolved bird tree of life, but they were used to answer pressing questions about brain evolution, bird's loss of teeth, and the evolution of song learning. As all of these genome sequences went public, eight papers were published in Science Magazine alone, and another 20 appeared in other journals. This work is changing the way that we view birds. And learning to deal with this much data is changing the way that we do science... Why does the order of birds in your favorite field guide change from edition to edition? Largely because ornithologists have been uncertain about the relationships among the major avian orders. Only recently have genetic data clearly shown that ducks and chickens (Galloanserae) make up a separate lineage from the rest of birds (Neoaves). But the biggest challenge in recovering the bird tree of life was that most bird orders seem to have appeared in a burst of evolution shortly after the other dinosaurs went extinct. It is almost as if they radiated into diverse forms to fill the void left by the departing dinosaurs. So bird researchers have known that it would take a lot of data - no, I mean A LOT of data - to resolve the relationships among bird orders and accurately date the origins of each order. But as several bird research groups were sequencing genomes of their favorite species to study things like neurological development, song learning, After several years of work assembling complete genomes from all of the major bird orders, researchers from around the world have published a suite of paper summarizing their analyses of bird evolution. Some of the cool insights: First let’s look at the phylogeny, which is a picture of the tree of life. Figure 1 from Science Magazine. Jarvis et al. 2014. Whole genome analyses resolve early branches in the tree of life of modern birds. Science 346(6215): 1320-1331.
This particular tree shows the branches as they occur, with time going from left (older) to right (younger), with a time-scale bar on the bottom of the tree. One obvious finding is that the major radiation of Neoaves bird orders occurred around the K-Pg boundary, 66 million years ago, around the time when the dinosaurs went extinct. Presumably the large void left when the dinosaurs and other animals went extinct made room for the surviving birds to diversify into the many land, sea, and air birds that exist today. The diversification happened rapidly, with most modern bird orders forming and diversifying by 50 million years ago. In the full genome tree, many of the traditional groups are retrieved in this phylogeny, but some aren’t. The falcons are split from the other hawks and eagles, and the falcons are related to parrots and songbirds. The tree also highlights many higher groupings of bird orders, including
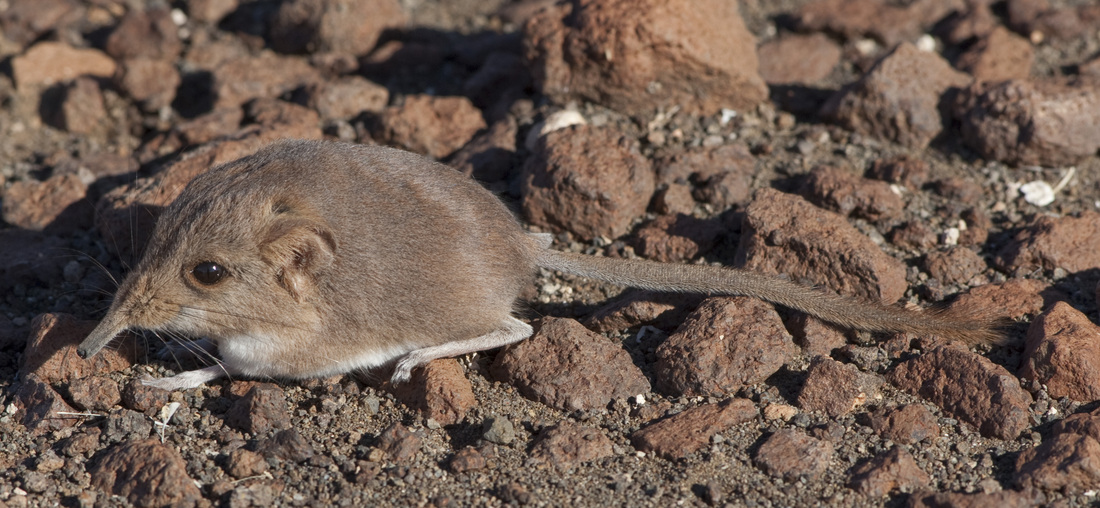
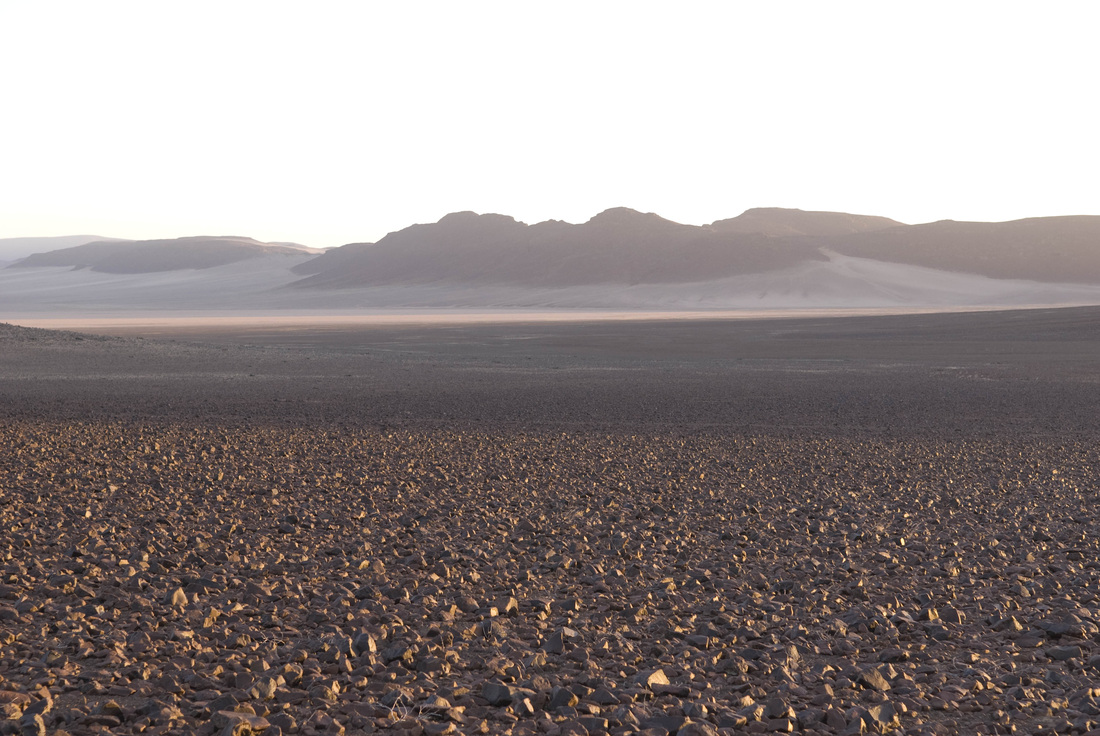
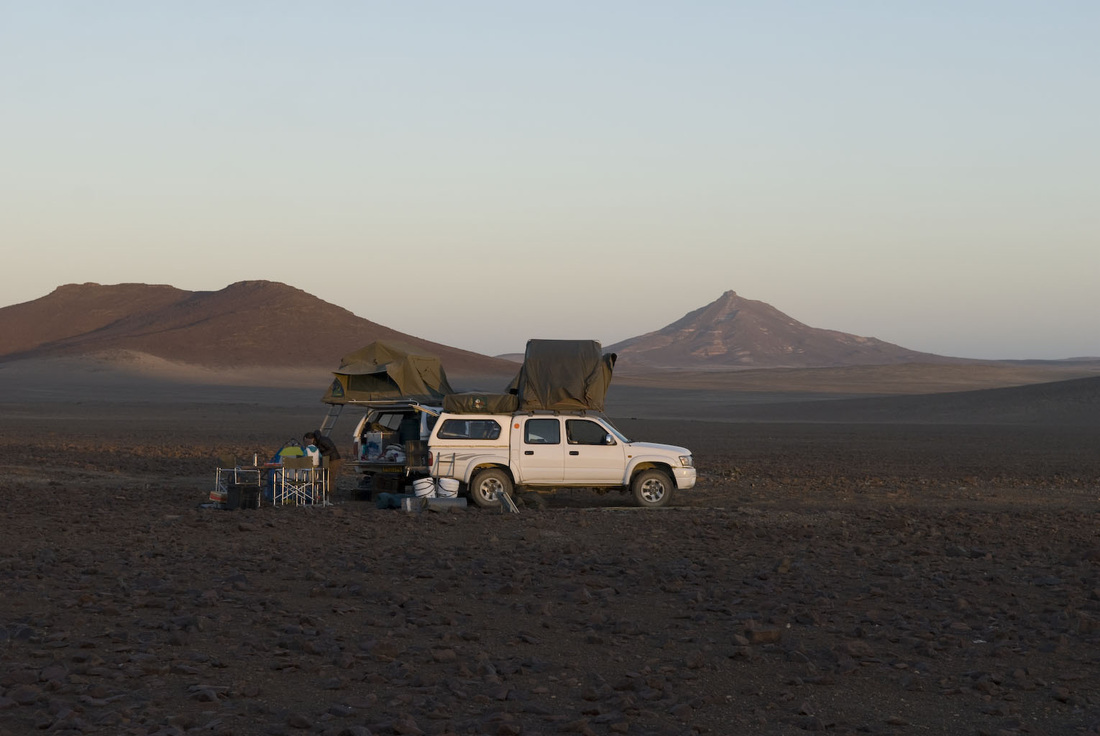
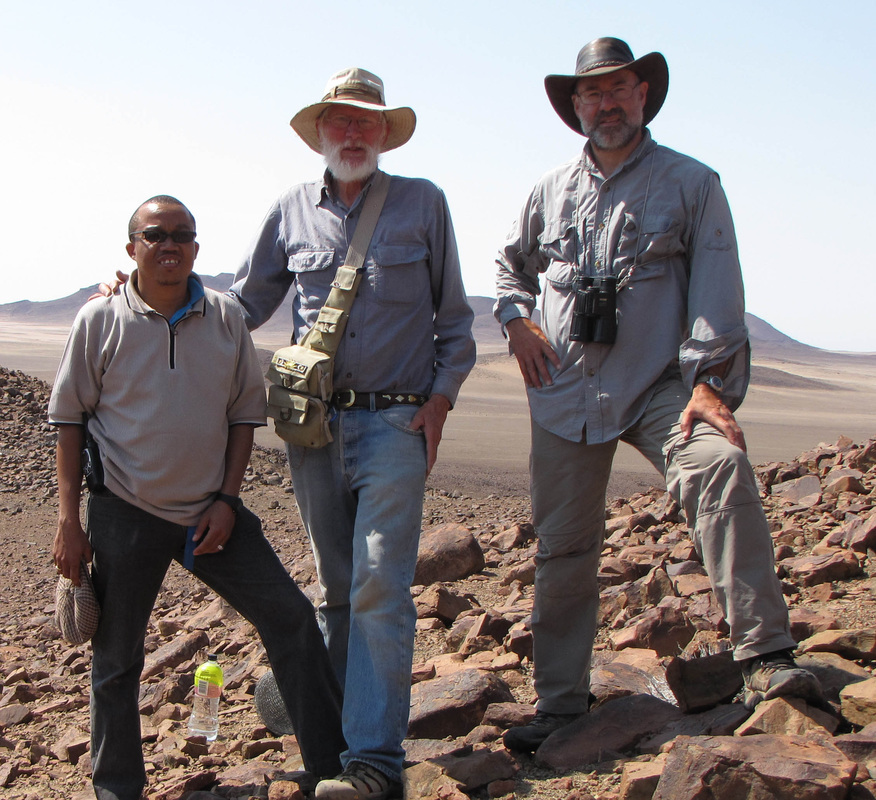
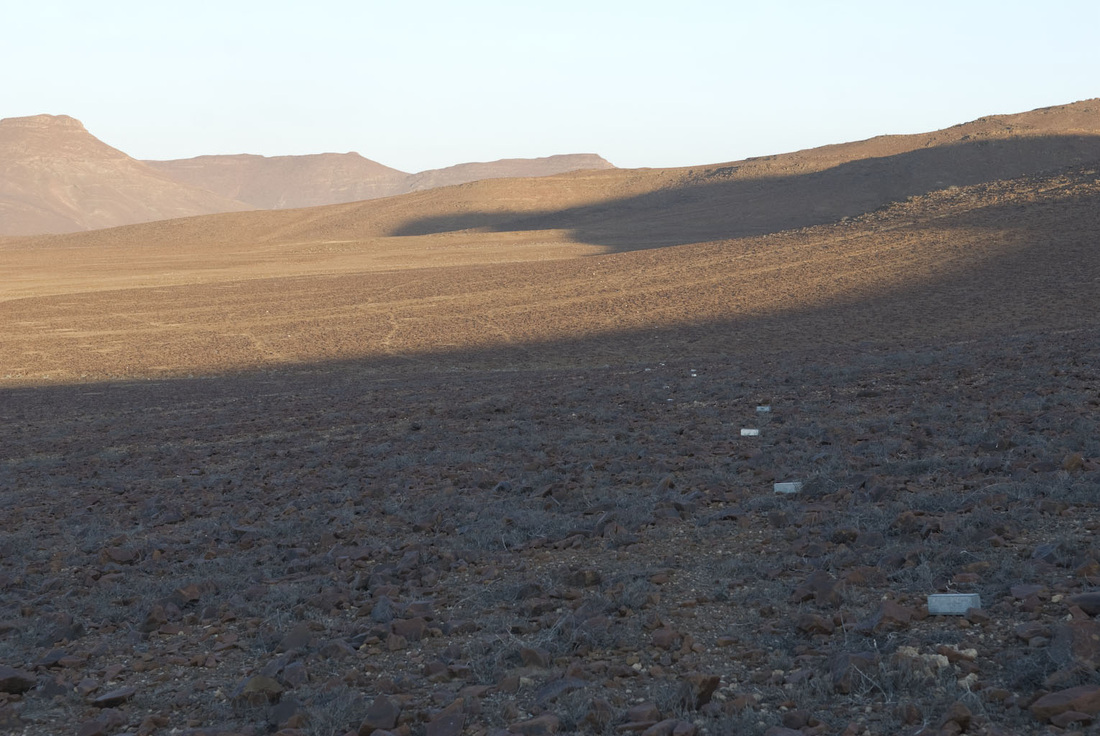
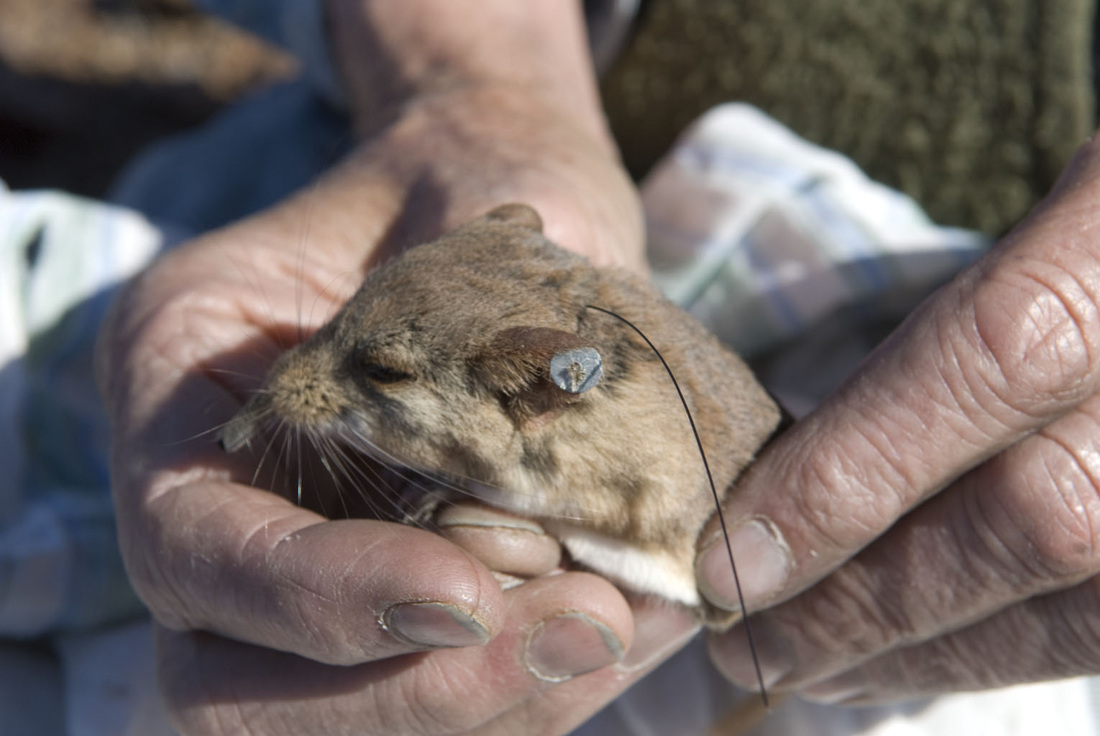
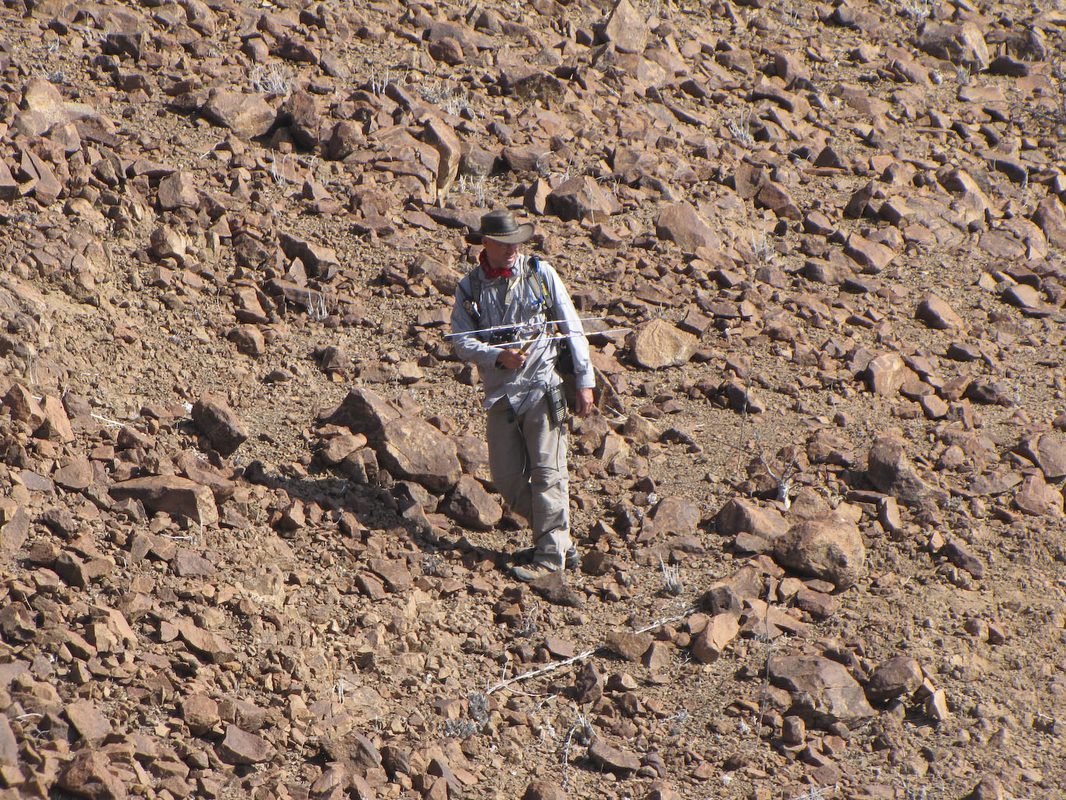
And finally, the order Passeriformes – which include nearly 50% of all living birds – started diversifying only about 40 million years ago, but has been one of the most rapidly evolving groups of birds. And there are several other genome findings, summarized in a second paper. Birds have smaller genomes than other non-fish vertebrates – and small genome size seems to be correlated with flight in general. For example, bats have smaller genomes than their terrestrial relatives, and typical birds have smaller genomes than their cousins who have lost the ability to fly (like ostrich.) Although this has been recognized and has sparked some interesting ideas, the genome data is suggesting that birds reduced their genomes by cutting out a lot of the junk – like transposable elements (aka jumping genes) and other repeat elements. Looking across the avian tree of life, it looks like most of the genome reduction happened very early in bird evolution. In fact, birds genomes are fragmented into many “microchromosomes”, and many of the missing pieces of DNA appear to be cut from the microchromosome break points. In other words, the smaller chromosomes appear to be cut up in ways that eliminated the junk between important genes. Most birds have a relatively similar arrangement of genes on their chromosomes, and across the avian tree of life, chromosomal rearrangements are relatively infrequent. It is unknown whether this is by design or simply by constraint. Most chromosome breaks and rearrangements are believed to occur at repeat regions – which are reduced in birds. It may be that by reducing the overall genome size, they have reduced their ability to make huge chromosomal rearrangements. This also means that there is a relatively slow rate of gene gain and gene loss in birds. Having full genomes across the avian tree of life allows researchers to more closely examine the role many genes throughout evolutionary history. For example, some traits, such as vocal learning evolved multiple times in birds, and the genes believed to be involved in vocal learning can be carefully studied to see how they change with respect to selection pressure. Likewise, the bird visual system, the oxygen carrying proteins in red blood, feather production can all be studied for evidence of natural selection and changes that control each of these systems. Virtually any scientist studying any bird trait can examine the evolution of genes involved in her favorite trait across the entire bird tree of life. These 48 genomes are just the beginning, and many more genomes are released each month. With this incredible amount of data, we can expect our knowledge of bird evolution to continue to take huge leaps forward. And maybe we can hope that the order of birds in our favorite field guides will become more standard and stable. My family and I just returned from 6 weeks of field work in Namibia, in southwestern Africa. The goal of the research was to study a small recently discovered sengi, also known as an elephant shrew... Earlier this year, my collaborators and I published a description of a new mammal species - a new sengi from the red-stone desert of Namibia. The species is the smallest member of the order Macroscelidea, or the elephant-shrews. Since they are neither elephants nor shrews, we prefer to call them "sengis." The new species was dubbed the Etendeka round-eared sengi, or Macroscelides micus. The common name refers to the Etendeka basalt region of northern Namibia where the sengi is found, and the specific name, micus, refers to its small size. This new sengi is small - weighing between 25 and 31 g (about 1 ounce) with a body of 85-95 cm long (about 3.5 inches long ). It's fur is redder than its relatives, probably to match the color of the red rocks where they live. Although they look a little like a mouse, their teeth and jaws are totally different from rodents. More like shrews, they eat mostly insects and arthropods. And they have a long moveable nose - probably suited to smelling and for lightly moving around soil and sand. Their habits are very poorly known. Limited data from relatives suggest that they probably have large home ranges, are mostly night-active, and probably monogamous. And this new species lives in some of the most barren landscapes I have ever seen, we were especially curious just how they can make a living here. And so we went to Namibia primarily to study their behavior in the wild. Based upon some work that Galen Rathbun did last year, we focused our field efforts on an area in the southern portion of the new species' range, just south of the Messum River and east of the large Brandberg Massif. This image shows the typical habitat, with ankle-high cobblestones strewn about rolling hills with sand settling in the lower plains and slopes. There are few low shrubs (mostly Welwitschia, Commiphora and Boschia), some grasses, and succulents. But in sum, there is very little plant life here. We made camp in rooftop tents because hyenas, leopards, and lions potentially roam the area. And there were definitely scorpions, spiders, and horned adders. And our cast of characters is here: Left to right are Seth Eiseb, Galen Rathbun, and me (Jack Dumbacher). Seth has worked with us on multiple projects, and was a curator at the National Museum of Namibia, but he is now a lecturer at the University of Namibia in Windhoek. Galen Rathbun is a research associate at the California Academy of Sciences, and a retired US Fish and Wildlife and US Geological Survey biologist. He is also the world expert on sengis, and has a great website on sengi biology. Also, not shown, is Tim Osborn, who is also a Field Associate of The Academy, and a long-time collaborator of Galen's and mine. To capture the sengis, we used small aluminum Sherman traps, about 9" x 3" x 3" placed on the ground about 10 meters apart. Sherman traps are little metal boxes with a door that shuts when a mammal goes inside. Rolled oats and peanut butter baited the sengis into the traps. We put out over 200 traps every night for over two weeks. After a few days, we captured one of the Etendeka round-eared sengis. We affixed a small radio collar around its neck (you can't see the collar, but you can see the small black antennae wire sticking up from the collar) and a reflective ear tag on its left ear. We also took weights and basic measurements. The animal was released near its trap site. During the field season, radios and ear tags were put on seven sengis. Using a small "H" shaped antennae and radio receiver, we were able to locate the direction of each sengi, and walk toward it until we found it. (Photo by Galen Rathbun) During the day, sengis sheltered under boulders or in holes under cobblestones. They were never active during the day. At night, the actively foraged, jumping from rock to rock, and looking for prey in crevices and under plants. The reflective ear tags helped us find them in the dark with just a headlamp and the radio receiver. When it was time to go, we caught all of the sengis by hand, and removed the ear tags and radio collars, and released them back into their shelters.
We are currently working up the data on home ranges, activity patterns, shelter characteristics, and hope to publish a short paper. Here are some other photos taken from the field site... |
Pages from the field
Jack Dumbacher's Blog. I am an evolutionary biologist and ecologist studying birds and mammals. I live for field work, but the genetics lab can be fun too... And living in the Bay Area is always full of surprises. Archives
November 2017
Categories |














































 RSS Feed
RSS Feed